This post may lead to a reduction in the amount of free time you have to drink tea or coffee while waiting for slow R code to finish. Read at your own risk.

Many will have heard the criticism from 'proper programmers' that R is slow. Just how valid is this criticism and when does it matter.
I recently submitted a beta version of my R package 'diveRsity' to CRAN. Since uploading the package, a few colleagues have tested it and by far the biggest issue they have encountered is its lack of speed. I knew before I uploaded the package that it wasn't particularly fast, but I was willing to live with what seemed like a reasonable analysis time of 3 hours for a data set of around 500 individuals (6 populations) genotyped for 37 microsatellite loci. The reasons why I could live with this snails pace were, 1) I was under the false impression that my data set was pretty big (if not in terms of the number of individuals, then the total number of genotypes.) and 2)I didn't now how I could speed up my code without using a different language or spending an excessive amount time learning proper R code and re-writing the package.
I attended a course last year in Seattle where, R core developer, Thomas Lumley said "Running code 2 or 3 times faster may not be worth the time spent [speeding up] code.... But going an order of magnitude faster is a good thing...." I wasn't sure if the effort required to learn vectorization and speed up my code would be worth the speed gains.
One thing I did know about my code was that it was full of loops within loops within loops within loops and loops are bad in R.
So, how to fix the problem? I first thought that I could learn to code R properly instead of the way that I had taught myself to. That seemed extreme, so I tried to 'parallelize' my code. Parallel computing is fast becoming the norm. Conceptually it is very simple. Divide multiple replications of the same job into groups and run these groups of jobs on separate processors/cores/CPUs, collate the results and hey presto your analysis is done.
Most modern PC and laptops have multicore processors making parallel computing possible, even at your desk.
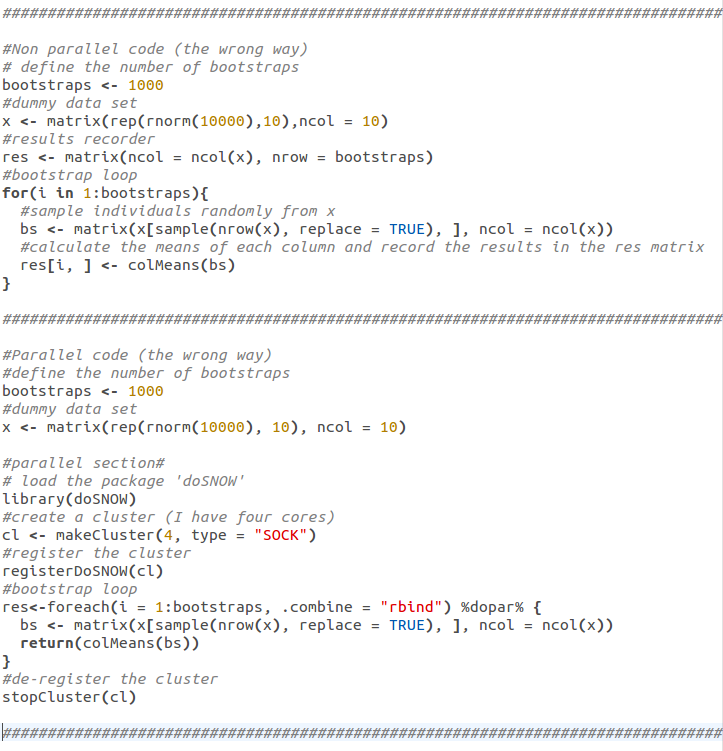
R has a plethora of packages which allow you to parallelize your code simply and without much decoding. See the code below which uses the 'doSNOW', 'foreach', 'snow' and 'iterators' packages for example. N.B. loading 'doSNOW' automatically calls the other three packages, that is why I've only called 'doSNOW' explicitly:
Parallelizing code:
By using a parallel approach to running my package code I gained about a 1/3 time reduction on Linux and a doubling in time in Windows!!!. Not as impressive as I had hoped. I finally realised that I was going to have to accept that I had written my code in R but had not written R code (i.e. vectorized code).
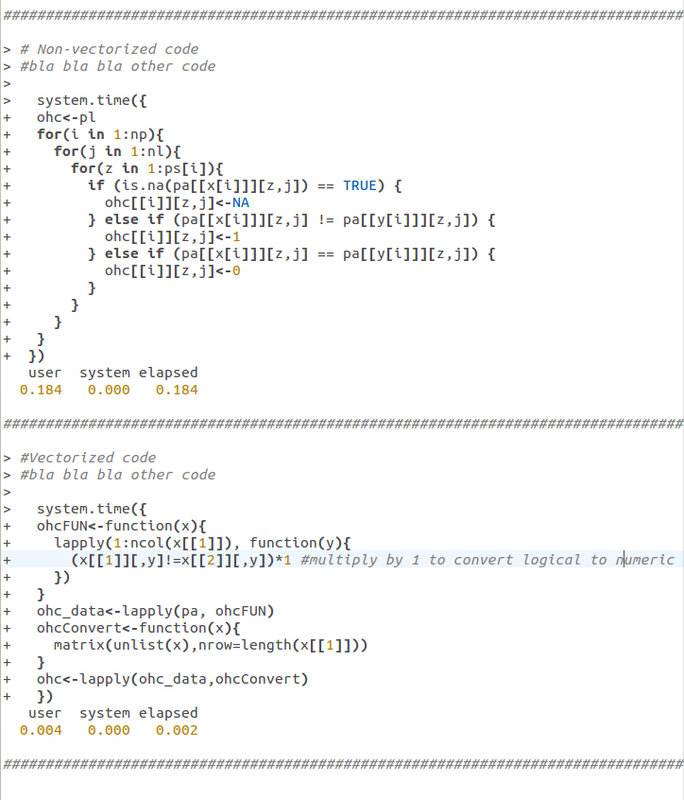
R, as a higher level language, has vectorization. It can apply a numerical method to groups of values much faster than it can apply the same numerical method to individual values within a group separately. As a simple example, here is some of the actual code from my package function 'pre.div' both before I vectorized it (top) and after (bottom):
Vectorizing code:
The time take to run the piece of code above will be:
(0.184 x 1000) x 15 = 2760 sec = 46 min.
For the vectorized code above only, the time taken will be:
(0.004 x 1000) x 15 = 60 sec = 1min.
Now 45min is time worth saving especially when you consider that this is the amount of time saved for a single chunk of code in a function containing approx. 20 similar chunks.
In summary, if you have slow code that you would like to be faster, first make sure that you have written R code and not just code in R, in other words vectorize where possible. Use functions like 'apply', 'lapply', 'mapply' etc. instead of 'loops', and if all else fails, go parallel.
Here are some links to resources that will get you started.
vectorization
The R inferno (see sections 3 and 4 specifically)
Parallel R
Parallel R loops in Windows and Linux
Parallel R book (N.B. purchase required)
 RSS Feed
RSS Feed
